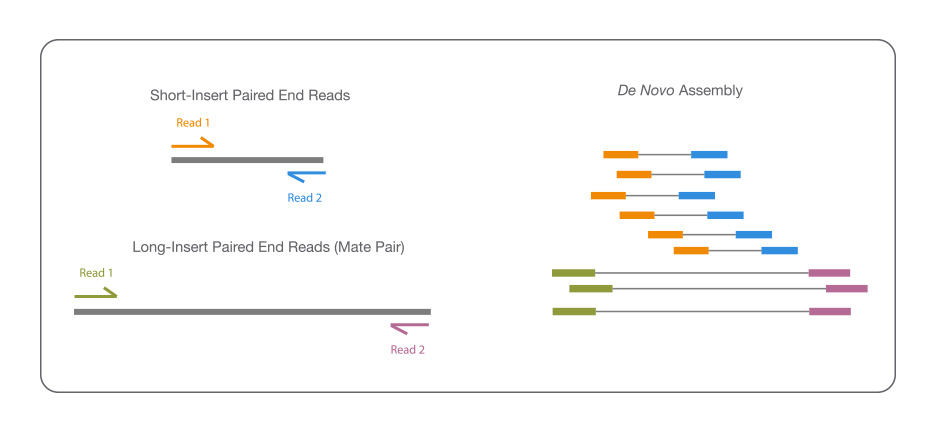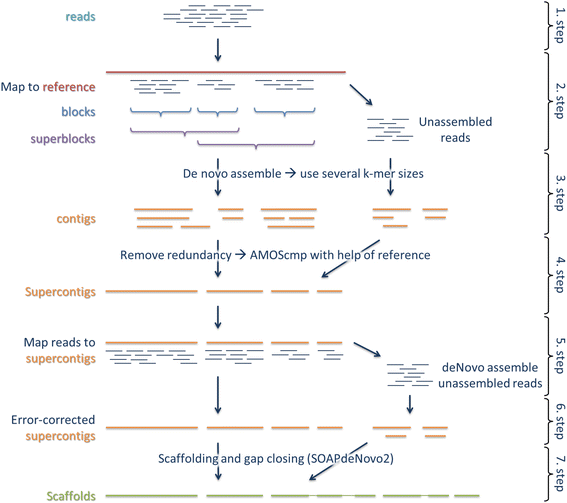
Reference-guided de novo assembly approach improves genome reconstruction for related species | BMC Bioinformatics | Full Text
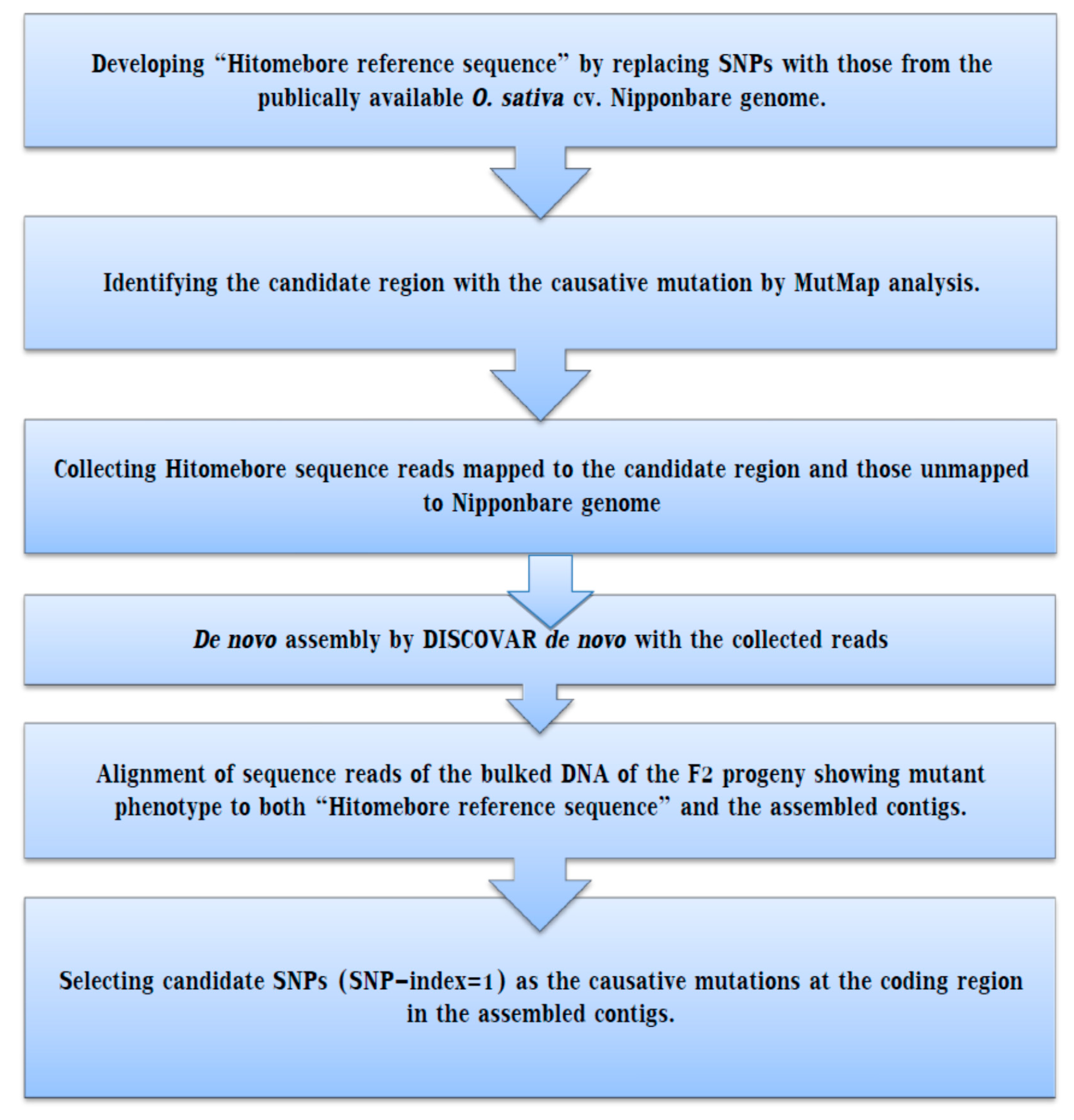
IJMS | Free Full-Text | Whole Genome Resequencing from Bulked Populations as a Rapid QTL and Gene Identification Method in Rice | HTML
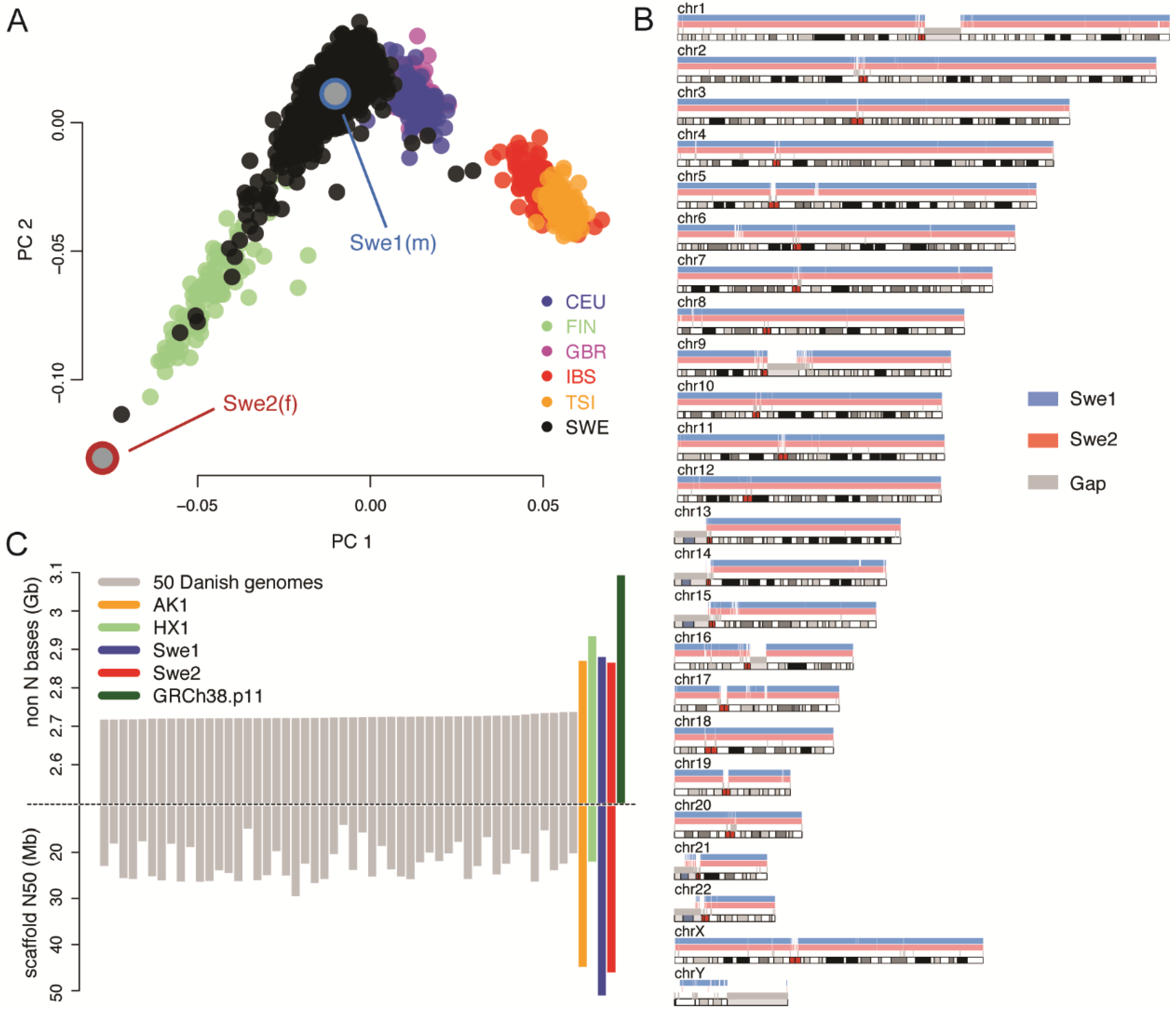
Genes | Free Full-Text | De Novo Assembly of Two Swedish Genomes Reveals Missing Segments from the Human GRCh38 Reference and Improves Variant Calling of Population-Scale Sequencing Data | HTML
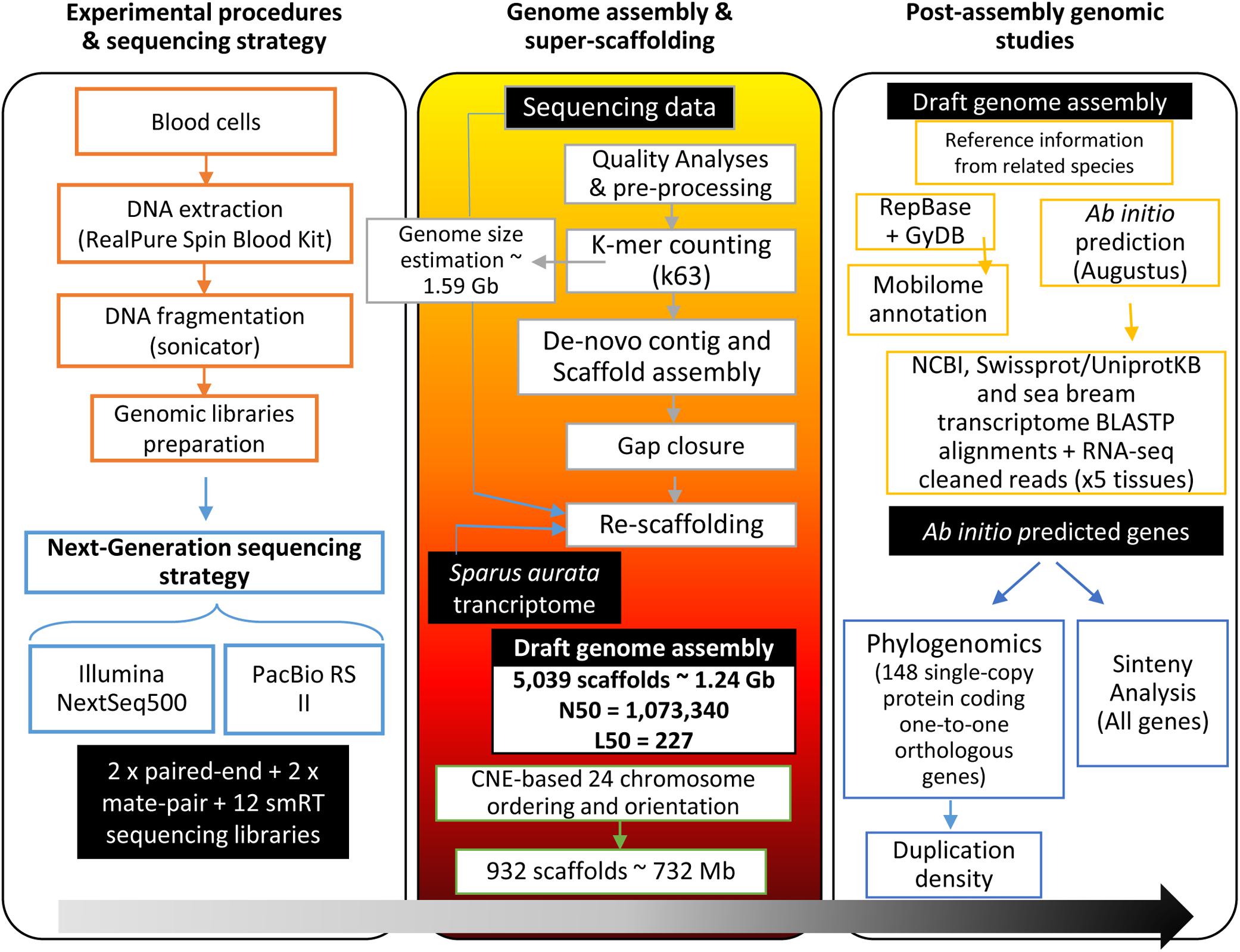
Frontiers | Genome Sequencing and Transcriptome Analysis Reveal Recent Species-Specific Gene Duplications in the Plastic Gilthead Sea Bream (Sparus aurata) | Marine Science

A comparative analysis of methods for de novo assembly of hymenopteran genomes using either haploid or diploid samples | Scientific Reports

De novo assembly of Oxford Nanopore MinION reads with QIAGEN CLC Genomics Workbench - Bioinformatics Software and Services | QIAGEN Digital Insights
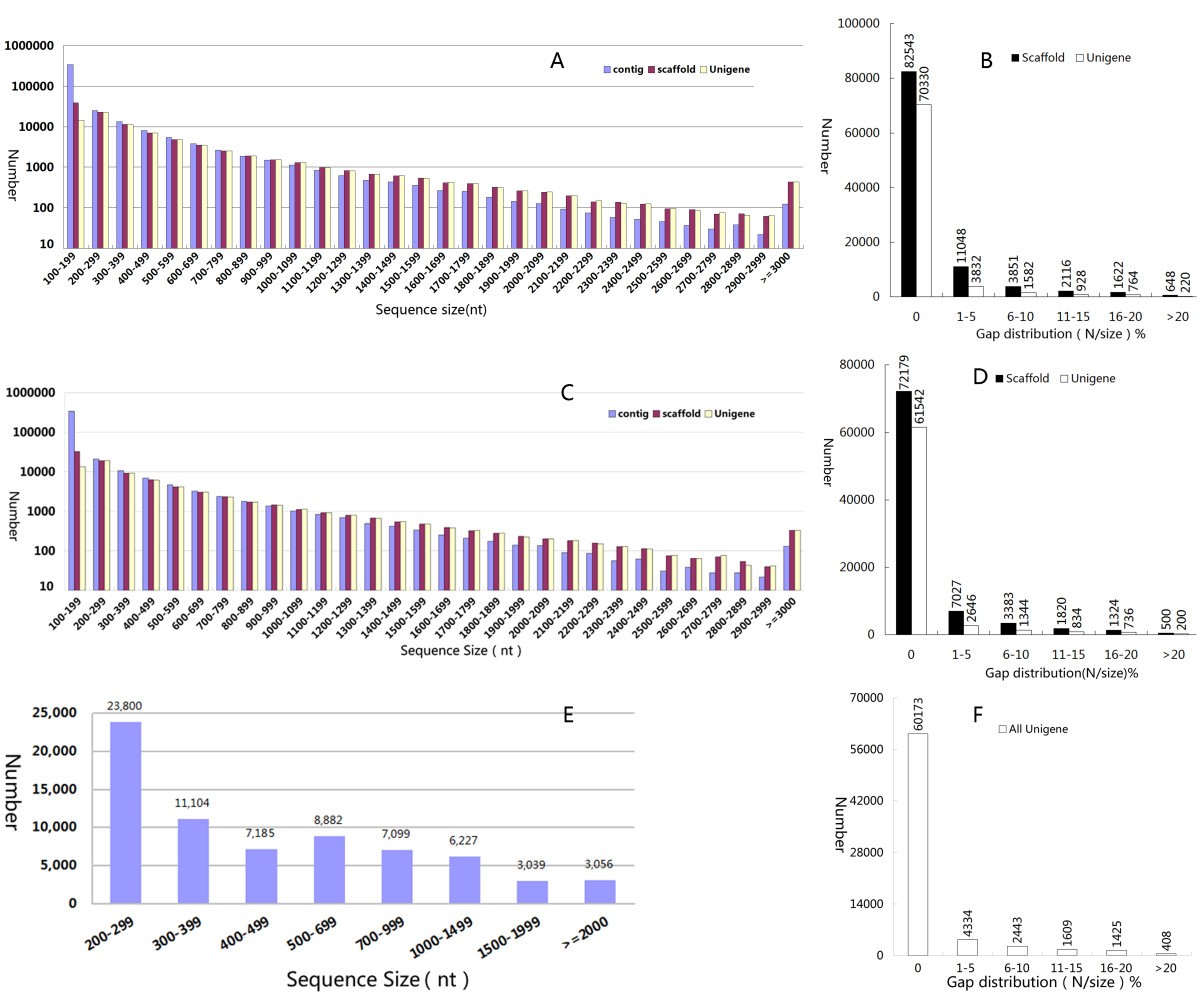
Transcriptome analysis of Sacha Inchi ( Plukenetia volubilis L . ) seeds at two developmental stages | BMC Genomics | Full Text
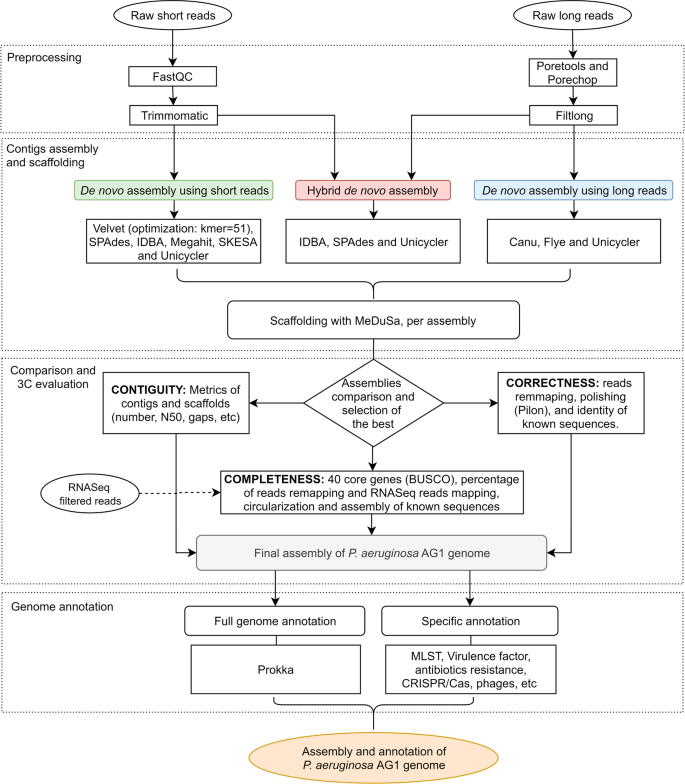
High quality 3C de novo assembly and annotation of a multidrug resistant ST-111 Pseudomonas aeruginosa genome: Benchmark of hybrid and non-hybrid assemblers | Scientific Reports
PLOS ONE: The Chaperonin-60 Universal Target Is a Barcode for Bacteria That Enables De Novo Assembly of Metagenomic Sequence Data
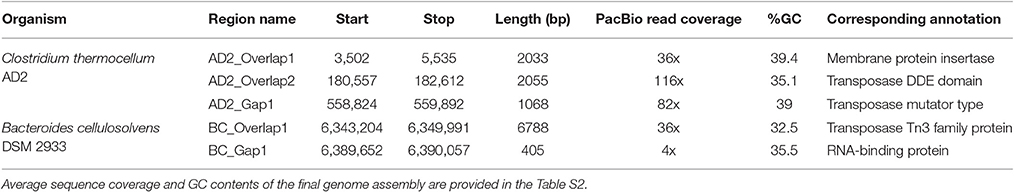
Frontiers | A Case Study into Microbial Genome Assembly Gap Sequences and Finishing Strategies | Microbiology







![Optimizing de novo genome assembly from PCR-amplified metagenomes [PeerJ] Optimizing de novo genome assembly from PCR-amplified metagenomes [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/6902/1/fig-2-full.png)